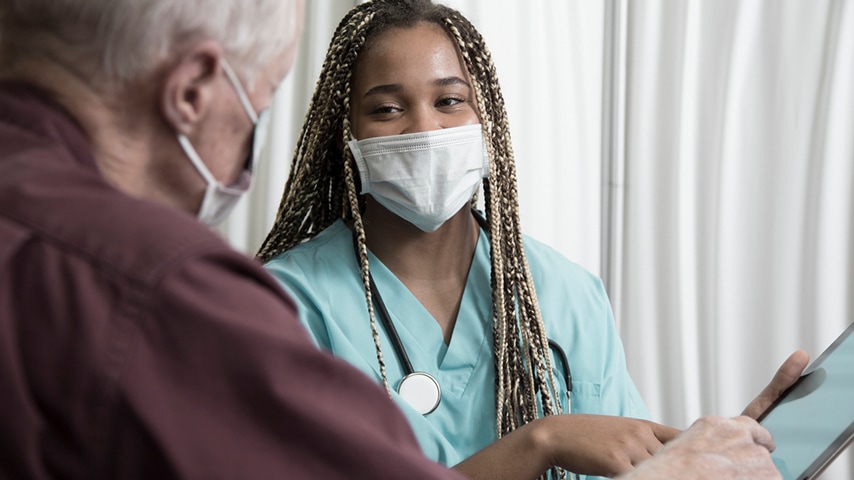
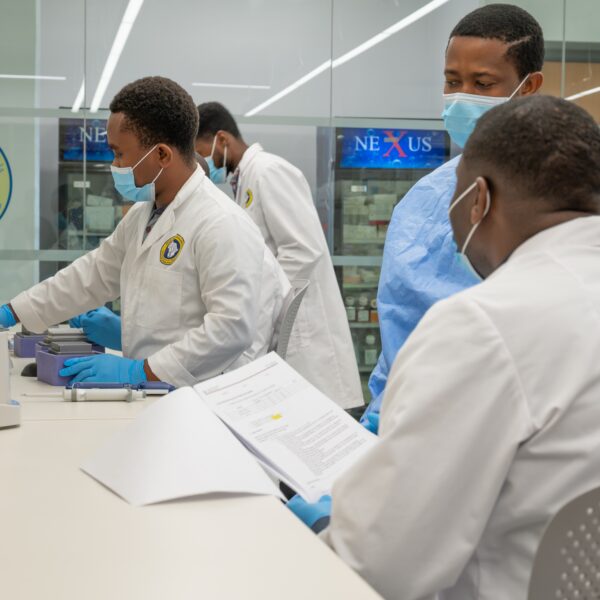
Diagnostics, Surveillance, and Epidemiology
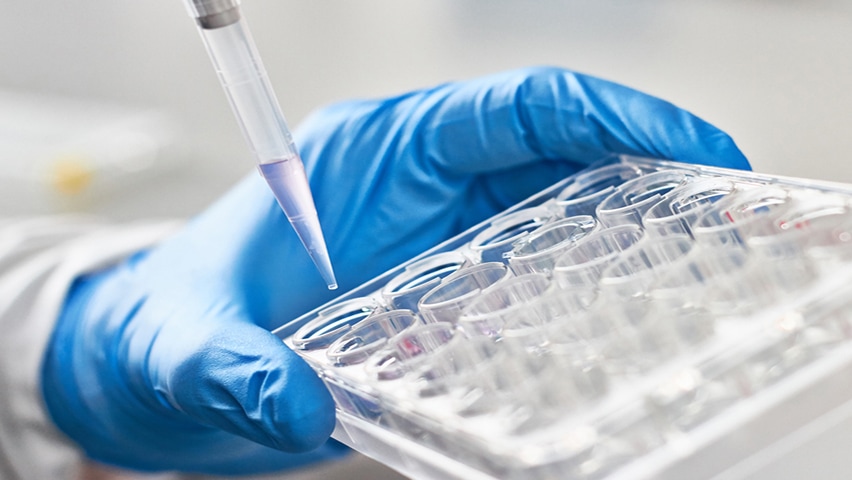

Our Focus and Working Group
The mission of the Diagnostics, Surveillance, and Epidemiology (DSE) Working Group is to drive collaboration and innovation in infectious disease detection, monitoring, and data-driven understanding. We aim to develop and implement improved diagnostics, surveillance tools, and epidemiological methods that strengthen both clinical practice and public health response.
Bringing together experts from academia, clinical medicine, public health, and industry, the group builds on lessons learned from COVID-19 to enhance our collective capacity to predict, detect, and prevent future infectious threats.
DSE collaborates closely with the New England Pathogen Genomics Center of Excellence (PGCoE)—a CDC-funded consortium —with the shared goal of advancing innovation and technical capacity in pathogen diagnostics, genomics, epidemiology, bioinformatics, and digital technologies, and accelerating their translation into public health practice.
Monthly presentations foster open exchange, collegial discussion, and new collaborations across disciplines.
Focus areas include:
- Novel detection technologies, including point-of-need, high-throughput, and multi-analyte testing
- Clinical and environmental surveillance (e.g., wastewater, air, and vector monitoring)
- Innovative serological approaches
- Tools for managing, analyzing, visualizing, and sharing data
- Public health applications of molecular and epidemiological modeling
- Decision-support tools for clinical and reference laboratories
Meetings: Third Friday of every month at 3:00 p.m. ET
Group Leads
-

John Connor, PhD
- Professor, Vice Chair of Virology, Immunology & Microbiology, Boston University National Emerging Infectious Diseases Laboratories (NEIDL)
-

David Walt, PhD
- Hansjörg Wyss Professor of Biologically Inspired Engineering, Harvard Medical School
- Professor of Pathology, Brigham and Women’s Hospital
- Core Faculty, Wyss Institute for Bioinspired Engineering at Harvard University
Latest Updates
Read MoreSupport Our Work
Help us stop infectious diseases and strengthen readiness for future outbreaks. As a non-profit, MassCPR relies on the generosity of donors to fund the research and clinical innovation that save lives.
More to Explore
Variants and Vaccines
Collaborate to track viral evolution and advance vaccine strategies
Post-Infectious Clinical Syndromes
Understand persistent symptoms, their underlying mechanisms, and impacts across populations





